-
Datasets are often accompanied by data dictionaries that define variables and data types so researchers can understand and reuse the data appropriately. MIDRC extends this concept by employing a graph-like, relational data model that explicitly defines relationships among different types of data. This approach supports FAIR principles, making data findable, accessible, interoperable, and reusable, and enables users to search across datasets, understand how data are connected, and combine information with confidence.
For example, MIDRC may contain records of medications taken by patients to treat a specific disease. To ensure these data can be interpreted correctly, the data model organizes them into related tables, or nodes, such as “cases” (patients) and “medications.” Each patient appears once in the cases node, while each medication record is linked to the corresponding patient, creating a many-to-one relationship. Similar links connect imaging studies, annotations, lab tests, diagnoses, and clinical encounters, allowing researchers to query complex clinical and imaging relationships without needing to manually reconstruct them.
A well-defined data model is also essential when combining MIDRC data with external resources such as the NCI Imaging Data Commons (IDC) or The Cancer Imaging Archive (TCIA). Precise definitions, often aligned with standard ontologies, support reliable data harmonization and reuse across platforms.
-
The MIDRC data model organizes structured data elements to support search across multimodal datasets in which data types may have complex relationships. It serves as the framework for data import, export, and harmonization, enabling data from disparate studies to be mapped to common standards. This allows users to identify comparable data across datasets and build cohorts within a unified web portal or through MIDRC Application Programming Interfaces (APIs).
Within the model, interrelated nodes (for example, “medication,” “imaging_study,” or “annotation_file”) contain category-specific properties such as medication name or body part examined. Records are connected through edges, which define links between data types, for example, associating imaging studies with patients or linking annotations to image files.
Technically, the model is implemented as a version-controlled JSON schema and it is maintained on the MIDRC GitHub and validated using Gen3 dictionary tools before release. Updates are proposed and reviewed by MIDRC subcommittees composed of radiology and data science experts, including the Data Standards and Information Technology (DSIT) and Data Quality and Harmonization (DQH) groups. Approved changes are tested in staging environments before deployment to production.
-
MIDRC primarily hosts images in DICOM format, except where DICOM files are unavailable. The imaging portion of the data model reflects the hierarchical structure of DICOM data (patient → imaging study → imaging series → image instances) and incorporates commonly used DICOM elements. Major imaging modalities, such as CR, CT, MR, MG, and ultrasound, are organized into dedicated nodes that link to imaging studies and ultimately to the patient record.
Learn more about DICOM:
https://dicom.innolitics.com/ciods
https://www.dicomlibrary.com/dicom/sop/ -
For clinical data associated with imaging, MIDRC drew on an established standard: the Observational Health Data Sciences and Informatics (OHDSI) OMOP Common Data Model. While adapted to accommodate MIDRC-specific requirements and the Gen3 framework, the core nodes and properties broadly align with OMOP, supporting consistency and interoperability with other research resources.
Learn more about OMOP:
https://ohdsi.github.io/CommonDataModel/
https://www.ohdsi.org/data-standardization/
https://athena.ohdsi.org/search-terms/start
MIDRC data model available on GitHub
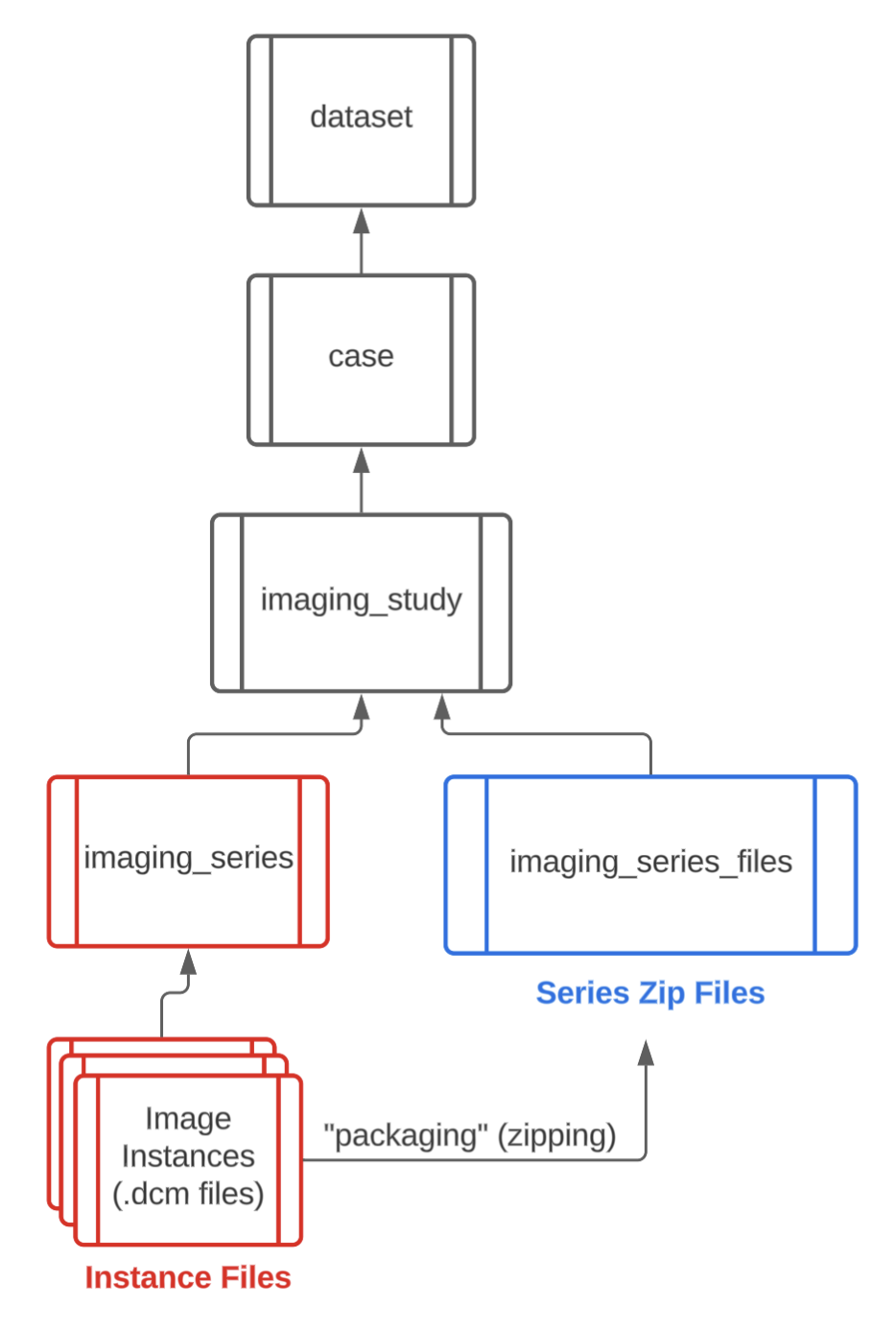
Core structure of MIDRC imaging nodes
Last updated January 30, 2026
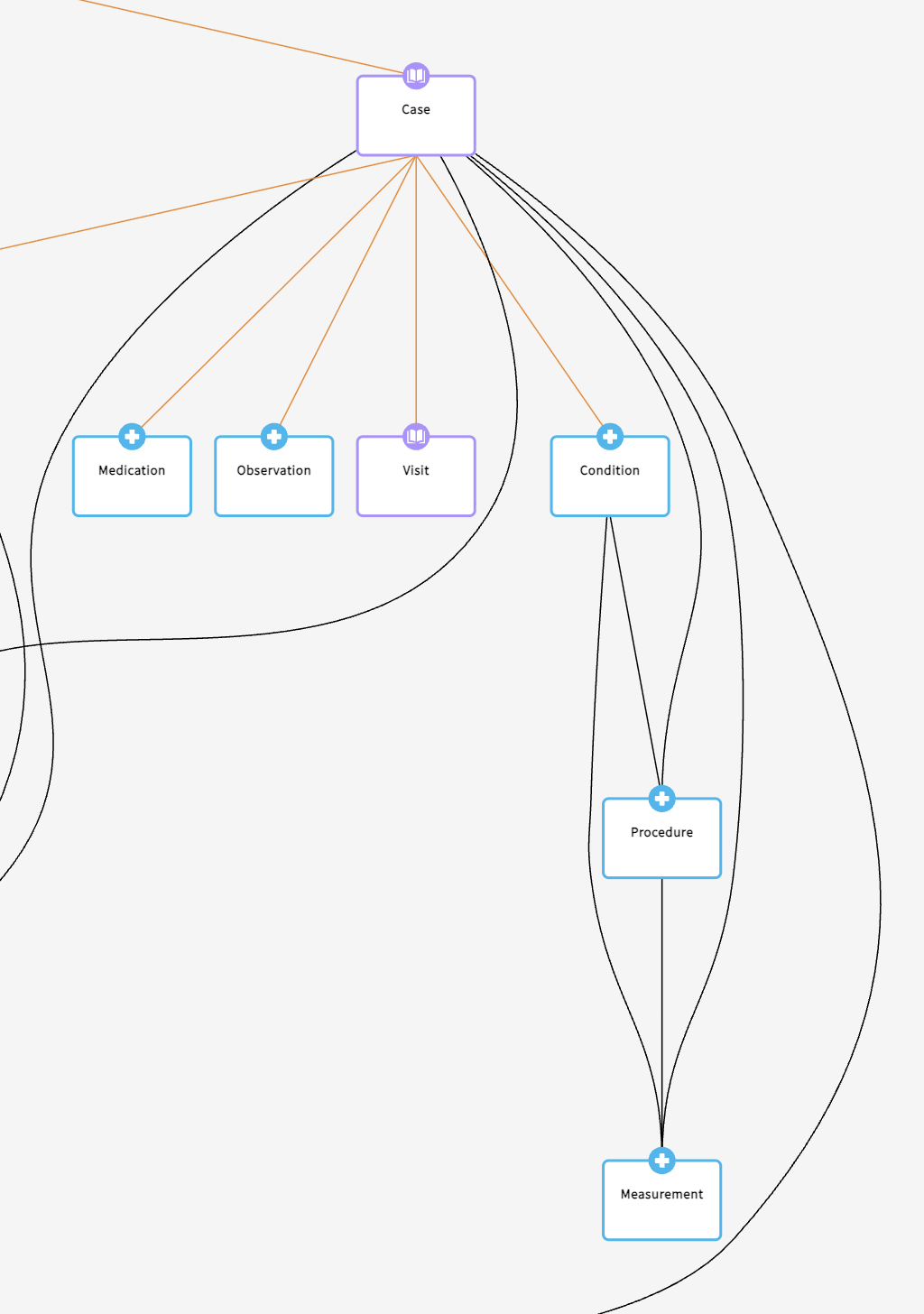
Core structure of clinical nodes
Data model.
Understanding MIDRC’s data model.

