Representativeness through interoperability.
This module is accompanied by an interactive Jupyter notebook.
The FAIR Guiding Principles for scientific data management and stewardship, published in Scientific Data in 2016, define four key principles—Findability, Accessibility, Interoperability, and Reusability—to enhance scientific data management.
Interoperability is defined as “the ability of data or tools from non-cooperating resources to integrate or work together with minimal effort.” According to the FAIR principles, for data to be interoperable, three requirements must be satisfied: 1) (meta)data use a formal, accessible, shared, and broadly applicable language for knowledge representation; 2) (meta)data use vocabularies that follow FAIR principles; and 3) (meta)data include qualified references to other (meta)data.
The importance of interoperability has been widely recognized in the healthcare community. Improved patient identification and data matching are critical for delivering high-quality and safe patient care. In this module, we emphasize the important aspect of interoperability—namely, how interoperability can improve data representativeness and accelerate the development and validation of artificial intelligence (AI) models.
There are two common scenarios for interoperability among data commons (Figure 1). Interoperability between data resources allows researchers to integrate multiple types of data from different data commons, including medical imaging, digital pathology, genomic, and clinical data (Scenario 1: multi-modal analysis). Interoperability also enables data from multiple institutions or cohorts to be combined with ease. The integrated data can then be used in subsequent stages of AI model development (Scenario 2: larger and more diverse datasets).
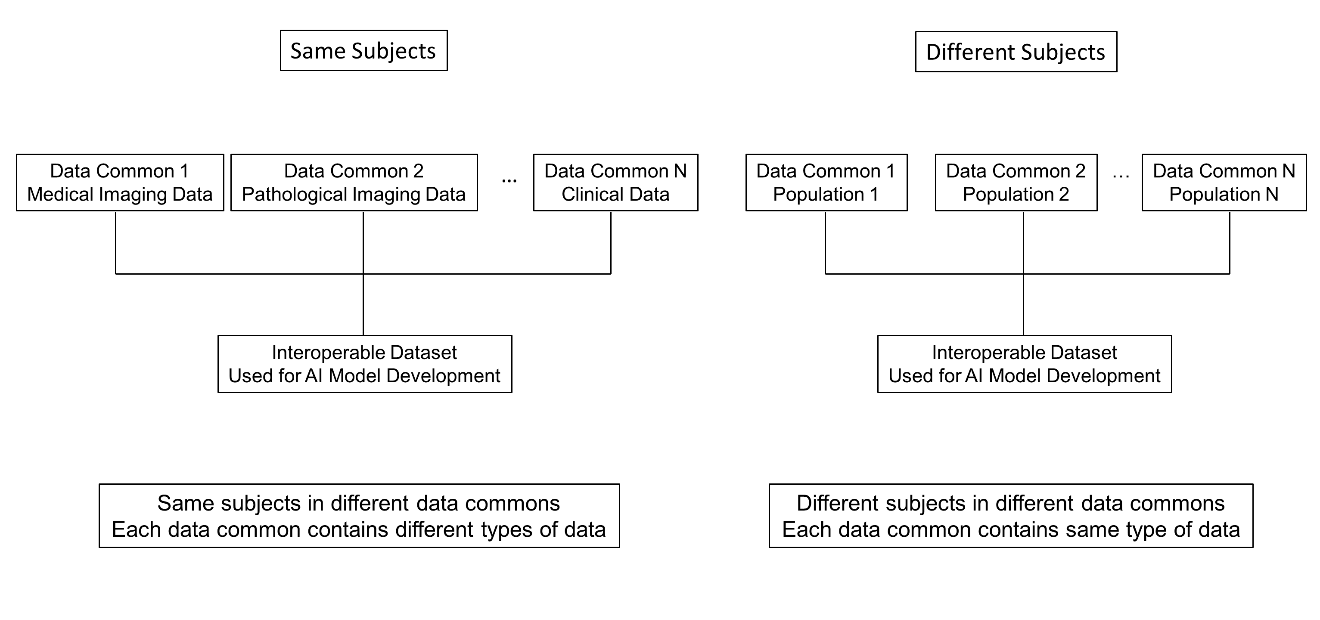
Fig 1. Interoperability among different data commons. (Left) The same subjects are represented across multiple data commons, and each data common contains a different type of data for these same subjects. (Right) Different subjects are represented across multiple data commons, and each data common contains the same type of data for these different subjects.
In Scenario 1, data from the same subjects are stored across multiple data commons, with each data common hosting a different type of data. For example, Data Common 1 may store medical imaging data, Data Common 2 pathological imaging data, Data Common 3 genomic data, and Data Common 4 clinical data for the same group of subjects. Because these data commons are interoperable, different data types from the same subjects can be integrated across commons.
The availability of comprehensive, subject-level data enables improved stratified sampling for AI model training and validation. For instance, because race and sex information can be obtained from the clinical data common, these attributes can be incorporated into the stratification strategy. As a result, subjects can be more evenly distributed between the training and test sets according to the race and sex distributions of the overall dataset. This leads to a more representative training set and helps reduce data selection bias.
For the validation set, inclusion of all race and sex subgroups allows model performance to better reflect the underlying population, thereby improving the fairness of model evaluation. In addition, the integration of data from multiple data commons enables true multi-modal analysis (e.g., radiomics–clinical–genomics analyses).
In Scenario 2, different subjects are stored in different data commons, and each data common contains the same type of data collected from different subjects. Interoperability—enabled through common data formats and shared vocabularies—allows data from multiple institutions and data commons to be used together in a consistent manner. Because these datasets follow common standards (e.g., DICOM and harmonized variable definitions), data from multiple institutions or data commons can be readily combined.
This interoperability enables the construction of larger and more diverse training datasets, increases demographic and scanner/protocol diversity, and reduces the risk of model overfitting to a single site. As a result, models are trained on more robust and clinically realistic populations, and the training dataset becomes more representative by including a larger and more diverse population.
For validation, interoperability enables truly external and site-independent validation. Model performance can therefore be evaluated across heterogeneous populations and institutions, allowing the reported results to better reflect real-world performance and improving the fairness of model evaluation.
Overall, interoperability enables standardized representation and exchange of multi-institutional data, facilitates scalable data integration, supports rigorous external validation, and improves the robustness and generalizability of AI model training and evaluation.
This module is accompanied by an interactive Jupyter notebook.

Last updated March 26, 2026
